Guide
This guide covers everything you need to use the gnomAD Carrier Frequency Calculator effectively. Whether you are a researcher or geneticist, these pages will walk you through the calculator's features and help you interpret results.
For Research Use Only
The gnomAD Carrier Frequency Calculator is a research tool. It is not a validated clinical diagnostic tool. Results must be independently reviewed and verified by qualified professionals before any use in a clinical context.
What is the gnomAD Carrier Frequency Calculator?
The gnomAD Carrier Frequency Calculator is a browser-based research tool that queries the Genome Aggregation Database (gnomAD) to calculate carrier frequencies for autosomal recessive conditions. It is designed for researchers and geneticists who need to explore population-level carrier frequencies and generate supporting documentation.
The calculator applies Hardy-Weinberg equilibrium to derive carrier frequencies from allele frequencies observed in gnomAD populations. Hardy-Weinberg equilibrium is a population genetics principle stating that, in a stable population, allele and genotype frequencies remain constant across generations. This means that if the pathogenic allele frequency in the population is q, then the carrier frequency (heterozygous individuals) is approximately 2_q_ for rare variants. The calculator uses this relationship to translate gnomAD allele data into interpretable carrier frequencies.
The tool supports multiple gnomAD versions - v4.1 (GRCh38), v3.1.2 (GRCh38, genomes only), and v2.1.1 (GRCh37) - and produces German and English documentation text. No account or installation is required; the calculator runs entirely in your browser.
Key Features
- gnomAD integration - queries multiple gnomAD versions (v4.1, v3.1.2, v2.1.1) directly from your browser
- Variant filtering - automatically filters for high-confidence loss-of-function (LoF HC) variants and ClinVar pathogenic/likely pathogenic entries
- Carrier frequency calculation - derives carrier frequencies from population allele data using Hardy-Weinberg equilibrium
- Recurrence risk estimation - calculates recurrence risk based on the index individual's genetic status
- Text generation - produces German and English documentation text with configurable perspective and gender-inclusive language styles
- URL sharing - export results as a shareable link for colleagues or records
- Search history - recent gene searches are saved locally for quick re-access
- Offline support - works as a Progressive Web App (PWA) after your first visit
The 4-Step Wizard
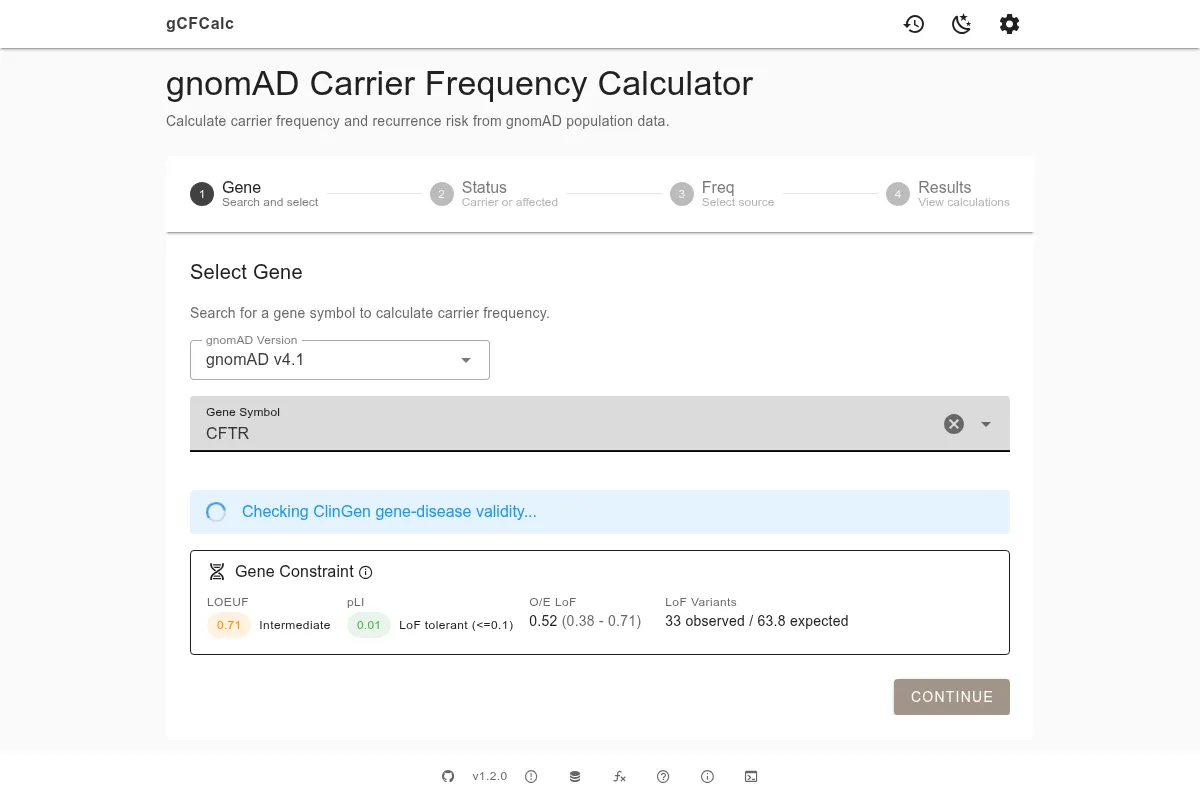
The calculator guides you through four steps from gene selection to text generation:
- Gene Search - find your gene by symbol and review gene constraint data to assess variant pathogenicity
- Patient Status - set the index individual's genetic status (heterozygous carrier, homozygous affected, or compound heterozygous)
- Frequency Source - choose between live gnomAD data, a literature value, or a default fallback frequency
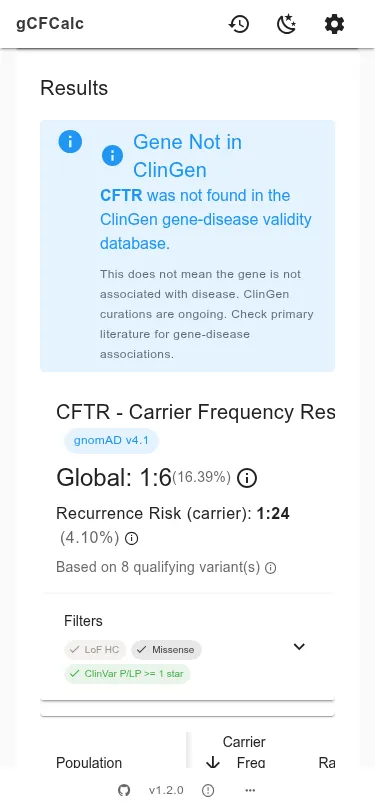
- Results and Text - review carrier frequencies by population, recurrence risk, and generate documentation text
Get Started
Open Calculator | Getting Started Guide
Next Steps
- Getting Started - a step-by-step walkthrough of a complete calculation
- Use Cases - example scenarios with worked examples
- Reference - methodology, data sources, variant filters, and template details