Data Sources
The calculator integrates data from the Genome Aggregation Database (gnomAD), ClinVar, and ClinGen. This page explains what each source provides and how to choose between gnomAD versions.
gnomAD Versions
The calculator supports three gnomAD releases. gnomAD v4.1 is the default and recommended choice for most calculations.
| gnomAD v4.1 | gnomAD v3.1.2 | gnomAD v2.1.1 | |
|---|---|---|---|
| Reference genome | GRCh38 | GRCh38 | GRCh37 |
| Data types | Exomes + Genomes | Genomes only | Exomes + Genomes |
| Samples | ~807,162 | ~76,156 | ~141,456 |
| Default | Yes | No | No |
| Unique population | Middle Eastern (mid) | Amish (ami) | Other (oth) |
When to Use Each Version
- v4.1 (recommended): The latest and largest dataset. Use this for most calculations. It includes both exome and genome data and covers the widest sample size across the most diverse ancestries.
- v3.1.2: Genome-only data on GRCh38. Use this when you need the Amish (
ami) population, which is not available in v4.1. - v2.1.1: Based on the older GRCh37 reference genome. Use for legacy comparisons or when working with variants annotated on GRCh37.
Population Codes
The three gnomAD versions cover overlapping but not identical population sets. Each population has a stable two- to three-letter code used internally by the calculator.
| Population | Code | v4.1 | v3.1.2 | v2.1.1 |
|---|---|---|---|---|
| African/African American | afr | Yes | Yes | Yes |
| Admixed American / Latino | amr | Yes | Yes | Yes |
| Ashkenazi Jewish | asj | Yes | Yes | Yes |
| East Asian | eas | Yes | Yes | Yes |
| Finnish | fin | Yes | Yes | Yes |
| Middle Eastern | mid | Yes | No | No |
| Non-Finnish European | nfe | Yes | Yes | Yes |
| South Asian | sas | Yes | Yes | Yes |
| Amish | ami | No | Yes | No |
| Other | oth | No | No | Yes |
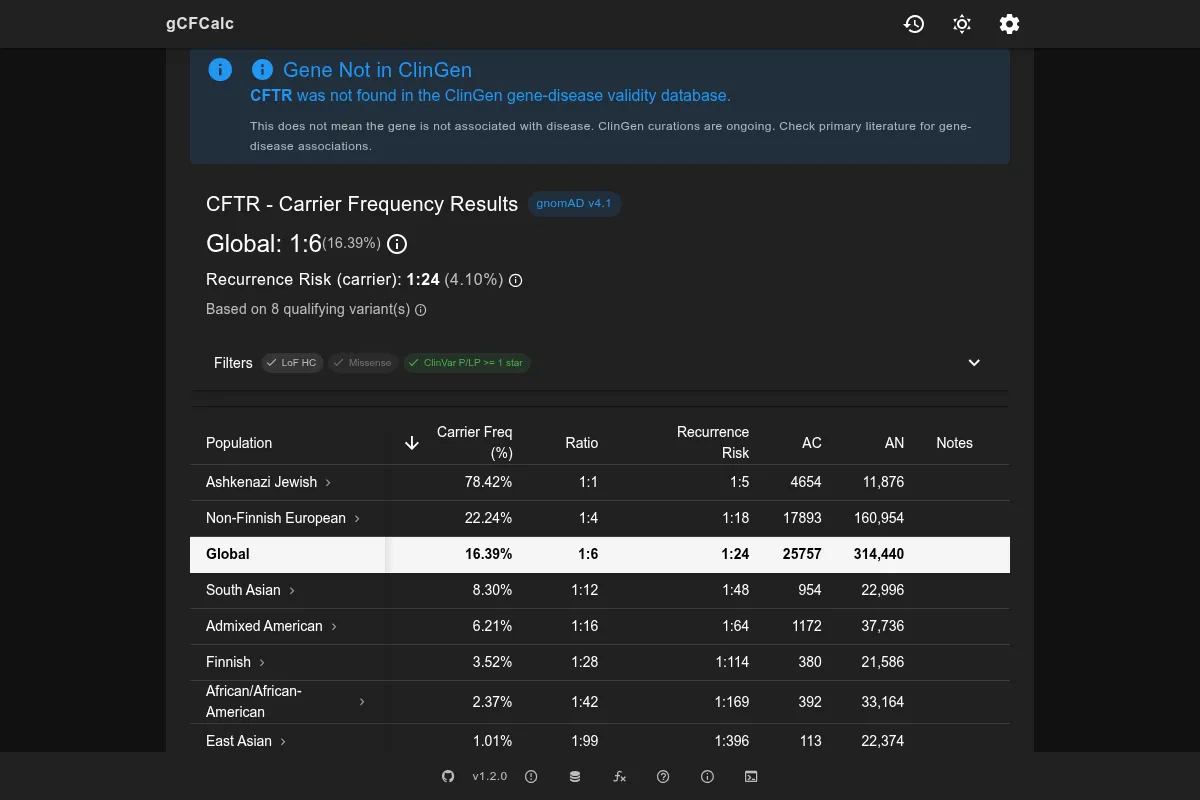
ClinVar Classifications
ClinVar is the NCBI database of genomic variants and their clinical significance. The calculator uses ClinVar classifications to determine which missense and other non-LoF variants to include.
- Classifications used: Pathogenic (P) and Likely Pathogenic (LP)
- Star rating: A 0–4 scale reflecting the level of review confidence
| Stars | Meaning |
|---|---|
| 0 | No assertion criteria provided |
| 1 | Criteria provided, single submitter (also applies to conflicting classifications) |
| 2 | Criteria provided, multiple submitters, no conflicts |
| 3 | Reviewed by expert panel |
| 4 | Practice guideline |
The default minimum star threshold is 2 stars. You can raise or lower this threshold in the filter settings. See Filters for configuration details.
ClinGen Gene-Disease Validity
ClinGen curates evidence for gene-disease relationships and assigns clinical validity classifications ranging from Definitive to Disputed. When a ClinGen classification is available for the searched gene, the calculator displays it as an advisory banner.
ClinGen is Advisory
ClinGen validity status helps you assess confidence in the gene-disease relationship, but the calculator does not filter variants based on ClinGen status. All variant inclusion is controlled through the Filters settings.
External Resources
- gnomAD Browser - Explore variant data directly in the gnomAD interface
- ClinVar - Search and review variant clinical significance submissions
- ClinGen - Gene-disease validity curation and clinical interpretation resources
See Methodology for how allele frequencies from these sources are used in calculations. See Filters for how ClinVar classifications determine variant inclusion.