Variant Filters
The calculator automatically selects pathogenic variants from gnomAD using a set of configurable filters. This page explains each filter, its default setting, and how it affects your results.
How Filtering Works
When you search for a gene, the calculator queries gnomAD for all variants in that gene. Filters determine which of those variants are included in the carrier frequency calculation. You can adjust filters globally in the settings dialog, or override individual variants in the variant table during a calculation.
Filter Defaults
| Filter | Default | Description |
|---|---|---|
| Loss-of-function (LoF HC) | Enabled | LOFTEE high-confidence loss-of-function variants |
| Missense | Enabled | Missense, inframe insertion, inframe deletion |
| ClinVar Pathogenic/Likely Pathogenic | Enabled | ClinVar P/LP classifications |
| ClinVar star threshold | 2 stars | Minimum review confidence |
| Include conflicting | Disabled | Variants with mixed ClinVar classifications |
| Conflicting threshold | 80% | Required P/LP percentage for conflicting variants |
Loss-of-Function (LoF HC)
Loss-of-function variants are assessed by LOFTEE (Loss-Of-Function Transcript Effect Estimator), a tool integrated into the gnomAD annotation pipeline. The calculator includes variants that LOFTEE classifies as high-confidence (HC) on the canonical transcript.
Included variant types:
- Stop gained (nonsense)
- Frameshift (insertion or deletion)
- Splice donor and splice acceptor variants
Key point: LoF HC is independent of ClinVar. A variant can be included based solely on its LOFTEE annotation, regardless of whether it has a ClinVar classification. This reflects the principle that high-confidence loss-of-function variants on a disease-causing gene are likely pathogenic by mechanism.
LOFTEE's high-confidence filter excludes low-confidence LoF variants that may be rescued by downstream translation re-initiation or other events.
ClinVar Pathogenic/Likely Pathogenic
The calculator includes variants classified as Pathogenic (P) or Likely Pathogenic (LP) in ClinVar, subject to a minimum star threshold.
The star threshold controls review confidence:
| Stars | Meaning |
|---|---|
| 0 | No assertion criteria provided |
| 1 | Criteria provided, single submitter (also applies to conflicting classifications) |
| 2 | Criteria provided, multiple submitters, no conflicts |
| 3 | Reviewed by expert panel |
| 4 | Practice guideline |
The default threshold is 2 stars. Raising the threshold excludes variants with fewer independent submissions and increases confidence in included variants. Lowering it includes more variants but with less review evidence.
Missense Variants
Missense, inframe insertion, and inframe deletion variants are included only when they have ClinVar P/LP evidence.
Key point: Missense variants always require ClinVar P/LP classification. Unlike LoF HC variants, a missense variant cannot be included based on computational prediction alone - it must have a ClinVar Pathogenic or Likely Pathogenic classification meeting the star threshold. This prevents unsupported missense inclusion, which would inflate carrier frequency estimates.
LoF HC vs. Missense: Different Evidence Requirements
LoF HC variants are included based on LOFTEE annotation alone (mechanism-based evidence). Missense variants require ClinVar P/LP classification (clinical evidence). This distinction reflects the higher predictive confidence for loss-of-function variants compared to missense variants.
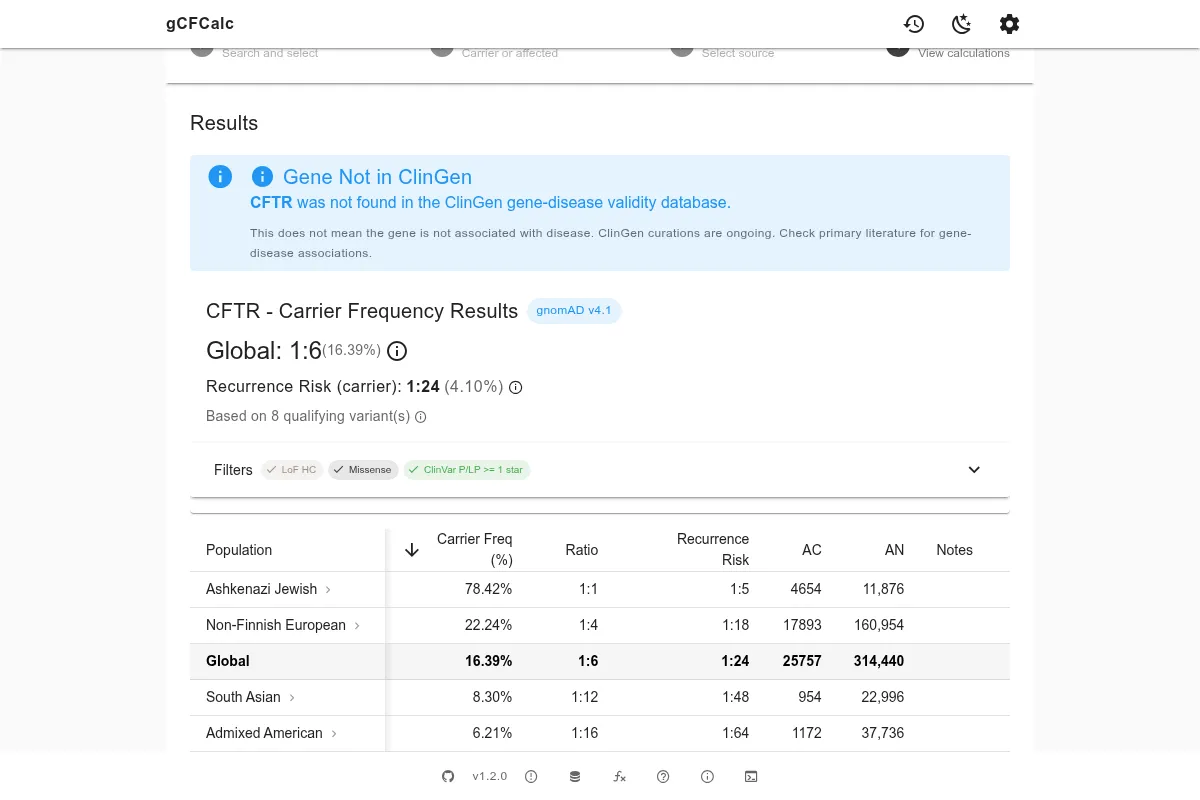
Conflicting Classifications
Some variants in ClinVar have submissions with mixed classifications - for example, some submitters classify a variant as Pathogenic while others classify it as a Variant of Uncertain Significance (VUS). These are listed in ClinVar as "conflicting classifications."
By default, conflicting variants are excluded. When you enable this option:
- Only conflicting variants where the P/LP percentage meets the configured threshold are included
- Default threshold: 80% - at least 80% of submissions must classify the variant as P/LP
- Example: A variant with 5 P/LP submissions and 1 VUS submission = 83% P/LP → included at the 80% threshold
Use Conflicting Classifications with Caution
Conflicting classifications indicate genuine uncertainty about pathogenicity. Enabling this option may include variants that are later reclassified. Review included conflicting variants individually in the variant table.
Per-Calculation Override
In the variant table (Step 3 of the wizard), you can manually include or exclude individual variants regardless of filter settings. This is useful for:
- Excluding a variant with disputed pathogenicity (for example, CFTR c.1210-11T>G, which has contested pathogenic significance)
- Including a variant you have independent clinical evidence for
- Investigating how a single variant affects the total carrier frequency
Manual overrides apply to the current calculation only - they do not change your global filter settings.
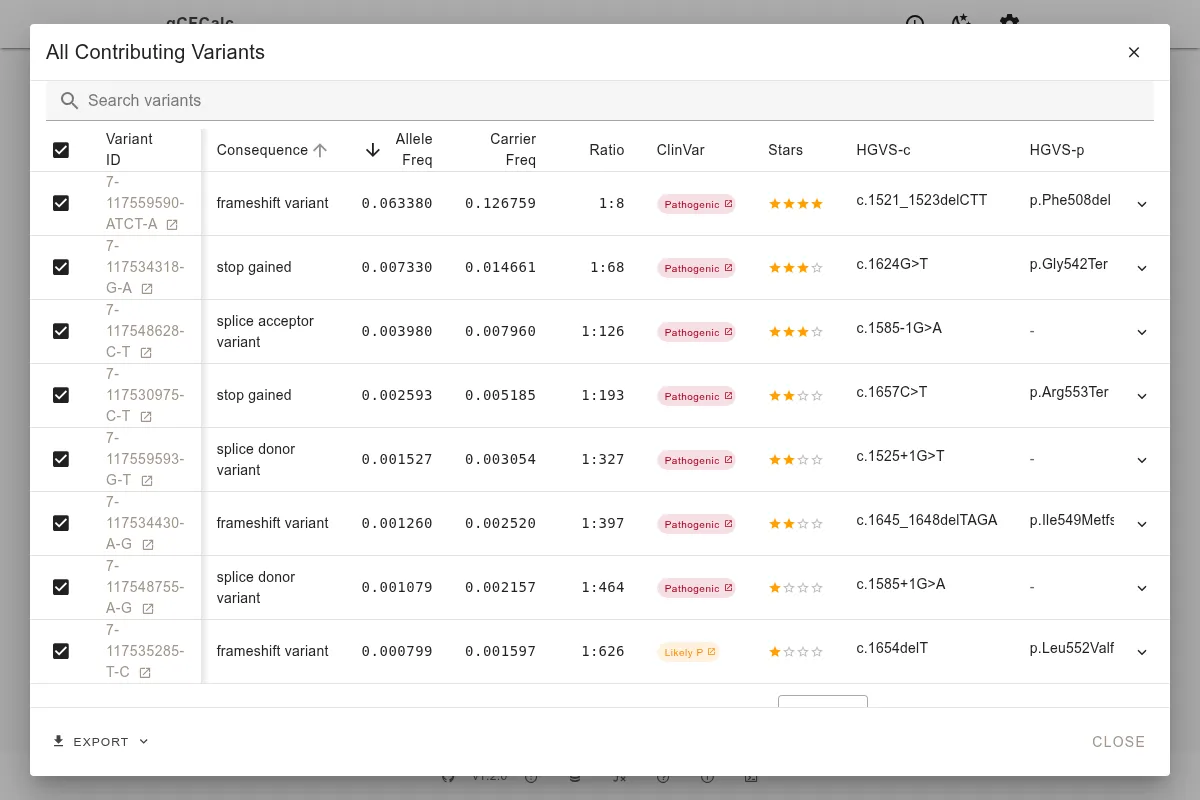
See Carrier Screening for a real example of manual variant exclusion with CFTR. See Methodology for how included variants are used in calculations. See Data Sources for ClinVar star rating details.