Carrier Screening Example
For Research Use Only
This example is for research and educational purposes. The calculator is not a validated clinical diagnostic tool. Any outputs must be independently verified by qualified professionals before use in a clinical context.
The Scenario
This hypothetical example demonstrates a carrier screening calculation using CFTR (cystic fibrosis). One individual is a known carrier of a pathogenic CFTR variant; the goal is to determine the carrier frequency in a given population to estimate recurrence risk.
This example covers arriving at a population-appropriate carrier frequency that accounts for the specific variants in the gnomAD dataset, and handling edge cases like disputed variant pathogenicity.
Using the Calculator
Searching for CFTR
Type "CFTR" in the gene search field and select it from the autocomplete list. The calculator queries gnomAD and retrieves all variants that meet the automatic inclusion criteria: Loss-of-Function High-Confidence (LoF HC) calls and ClinVar pathogenic/likely pathogenic classifications.
Reviewing the Variant Table
When the CFTR variant table loads, review each variant before accepting the defaults. The automatic filters capture the right variants for most situations, but CFTR is a gene where judgment on variant inclusion matters.
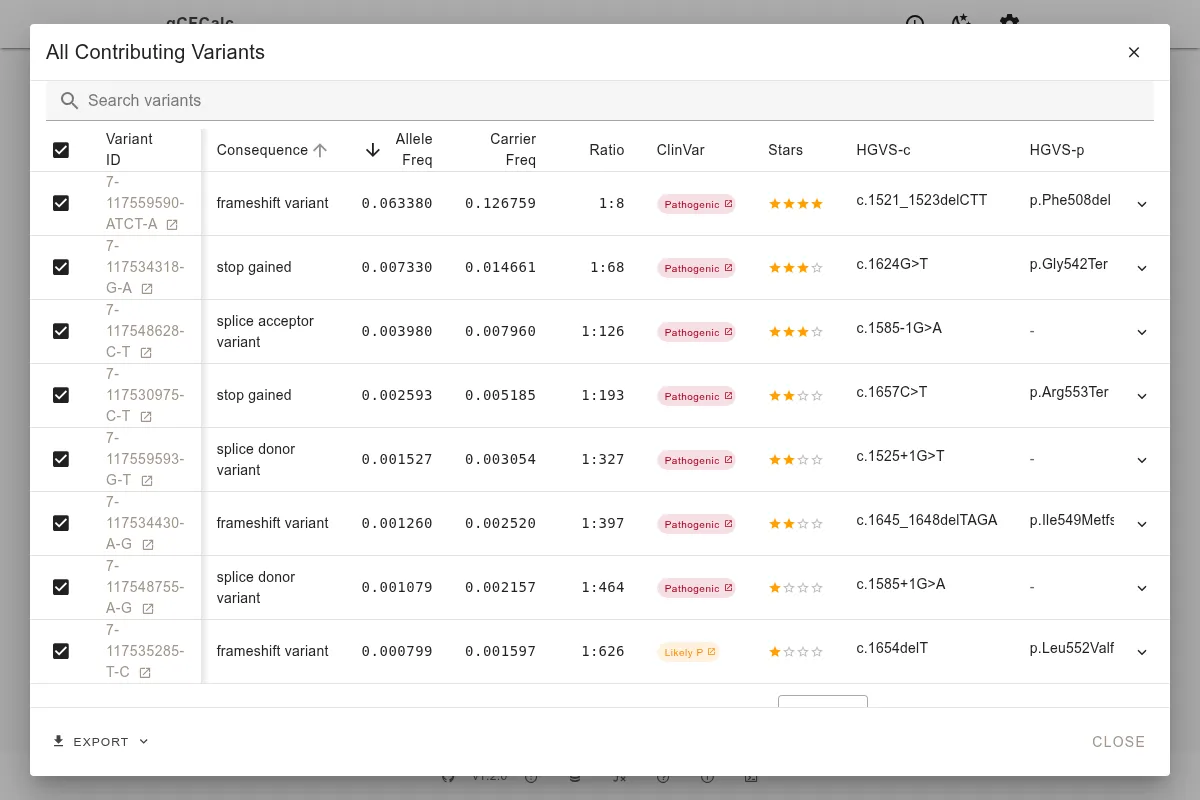
One variant that may appear is c.1210-11T>G - also known as the 5T allele in the intron 8 polypyrimidine tract. This variant has context-dependent significance. Its pathogenicity depends on the TG repeat tract length on the same allele (TG12 or TG13 increase severity) and whether a classic CF-causing variant is present on the other allele. Its presence in gnomAD at appreciable frequency reflects this complexity: it causes complete cystic fibrosis only in specific combinations, and isolated carriers may have no phenotype or only CBAVD (congenital bilateral absence of the vas deferens).
Excluding a Variant
To exclude c.1210-11T>G from the carrier frequency calculation, uncheck it in the variant table. The carrier frequency updates immediately using only the remaining variants.
This lets you compare the carrier frequency with and without the disputed variant and document the basis for your choice.
When to Exclude Variants
Variant exclusion is useful when a variant's pathogenicity is disputed or context-dependent. The calculator lets you see the impact on carrier frequency with and without specific variants. See the Filters reference for details on automatic variant filtering criteria.
Interpreting Results
The results table shows carrier frequencies for each gnomAD genetic ancestry group. Focus on the population that best matches the individual's ancestry - this is where the carrier frequency will be most relevant.
With heterozygous carrier status selected, the recurrence risk equals:
carrier frequency x 1/4
Because one individual is a known carrier, the remaining question is whether the other individual also carries a pathogenic variant (probability = population carrier frequency). If both are carriers, the probability of an affected offspring is 1/4 (standard autosomal recessive Mendelian risk). The calculator therefore computes: carrier frequency / 4.
ClinGen Classification is Advisory
ClinGen gene-disease validity classifications are shown in the results for reference but do not automatically filter variants. The variant table criteria (LoF HC + ClinVar pathogenic/likely pathogenic) are what determine the calculation inputs.
Next Steps
After reviewing the carrier frequency:
- Document which variants were included and the reason for any exclusions
- Note the gnomAD version and ancestry group used
- Use the text generation feature to produce documentation text
Try It Yourself
Ready to calculate carrier frequency? Open the calculator pre-loaded with CFTR data:
Or start fresh with any gene: Open Calculator
See Also
- Methodology - Complete explanation of the carrier frequency and recurrence risk formulas
- Filters - How variants are automatically selected for inclusion